Biochemical reactions
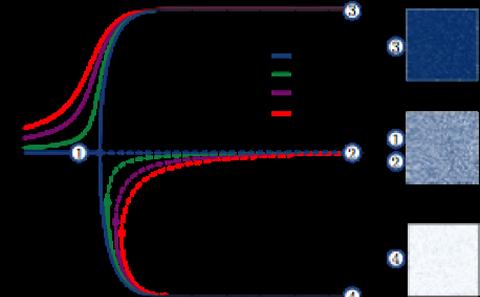
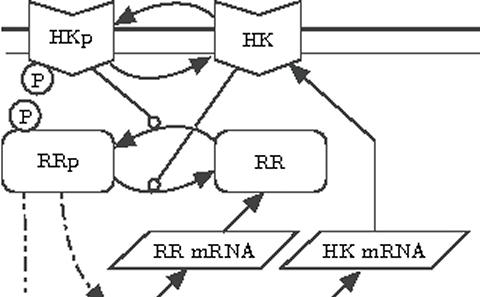
It is now well known that intrinsic noise can drive important biochemical processes. Intrinsic noise is associated with the uncertainty of knowing when a reaction occurs and what the reaction is. These effects are particularly accentuated when there are small numbers of molecules in the system. In this case the kinetics between the different species is best described by discrete-space continuous-time Markov processes. One of the main issues with this description is that even relatively small biological models very quickly become computationally intractable. Our main work in this area relates to the design and analysis of algorithms that are able to overcome this computational bottleneck.