Nuclear magnetic resonance is an extraordinarily powerful technique to not only solve molecular structure, but investigate molecular dynamics, ligand interaction and affinity, substrate turnover, metabolomics, membrane/protein effects and much more.
Nuclear magnetic resonance occurs when certain nuclei are placed in a magnetic field and irradiated with very weak electromagnetic radiation (radio waves). The nucleus type must possess a property called spin and those with spin=1/2 are the most commonly studied. these include 1H, 13C, 15N, 19F and 31P amongst others.
Biomolecules are often labelled with these nuclei (eg 12C replaced with 13C) so more complex experiments can be carried out and more information gleaned.
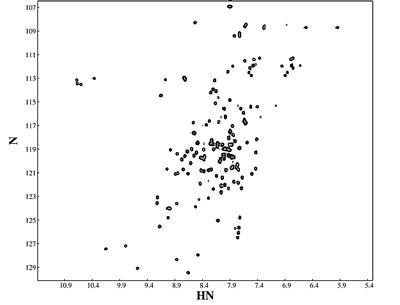
In solution NMR, the most common studies carried out at the Centre for Biological Sciences involve protein interactions and dynamics. Of all the multiple signals we see in the NMR spectrum we can follow a process called sequential assignment whereby each 1H, 15N and 13C in the protein is assigned to particular peaks in the spectrum.
Once assignment is complete, we can see which signals (and therefore which nuclei and which amino acids) are affected by the addition of ligands, drugs, peptides, DNA or other proteins. These can be used to map binding sites and measure affinities.
By changing how we apply the radio frequency pulses we can change the information reported back from the nuclei. NMR is the most advanced technique for investigating molecular motion and can provide dynamical information at many different timescales. Samples prepared in specialised alignment media can be studied to investigate
Chemists use NMR as their principal technique for identifying and confirming what they have synthesised in the laboratory. Drugs isolated from natural product extracts (eg medicinal plants) are characterised using a blend of techniques but NMR is the key method.
Solution NMR samples are typically 0.5mL or less in volume and are prepared in water or organic solvents but these must be (at least partially) deuterated eg D2O or CDCl3. The concentration required depends on the type of work to be carried out but ranges from 50μM to 1mM for protein assignment and structural work.
Solution NMR has the benefit of the moleculare tumbling freely (isotropically) in solution. This is not the case for NMR in the solid state and this has implications. Two main issues arise (chemical shift anisotropy and dipolar coupling between nuclei) that lead to severe broadening of the NMR signals so they can no longer be differentiated. Magic angle spinning (MAS) can reduce these and bring back the sharp signals by spinning the sample tube at high speed (10-100kHz) at an angle of ~54.7°. This rotationally superimposes the three dimensions of space removing the fixed orientation of nuclei in the solid sample so they appear isotropic. Dedicated probe technology using high pressure air drives the sample spinning.
Biological samples commonly studied using solid state NMR include membrane proteins and amyloid fibils such as those in Alzheimer's and Parkinson's disease. Samples are packed or centrifuged into an NMR rotor. Rotors vary in diameter from 6mm to 1.6mm. the smaller they are the faster they can spin. In biology, the common size is 3.2mm which holds 21μL of sample.
To make use of the NMR facility or enquire as to what it could achieve in your research, speak to Joern Werner in the first instance.