Child page cards
-

About us
An interdisciplinary research institute bringing together expertise from across the University to tackle society's challenges in the Life Sciences. -

Our people
Learn more about the Institute for Life Sciences management group and academics leading strategic interest groups. -

Annual report
Our annual report features high impact case studies of interdisciplinary Life Sciences research. -

Interdisciplinary health and medicine
We are interested in interdisciplinary science that addresses challenges in health and medicine. -

Interdisciplinary data insights
Our researchers at Southampton are developing cutting-edge mathematical and computational methods to advance data technologies. -

Interdisciplinary life technologies
Our interdisciplinary activity in Life technologies spans development of fundamental imaging technologies through to their application addressing key grand challenges in the life sciences. -

Interdisciplinary living systems
Research in this area addresses critical challenges ranging from the molecular scale to complex biological systems and environments. -

Highlights
Search our archive of interdisciplinary Life Sciences research news stories. -

Regional life sciences
The central south of England has a collaborative, research-active life sciences sector with strong academic, clinical and commercial links.
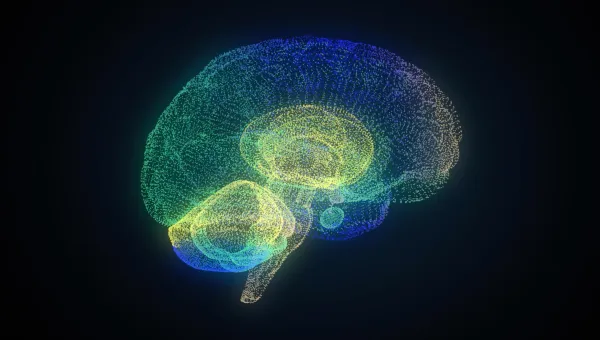
Pioneering Alzheimer’s drug development
Advancing the development of targeted treatments to slow or halt disease progression.
-

Transforming chronic respiratory disease care
-

Exploring how young people's diet, environment and mental wellbeing shape their health
Southampton researchers explored how food insecurity, diet and urban life shapes adolescent health in Soweto.
-

Dealing with chronic pain
-

Rapid diagnosis of dementia
A new rapid laser-based test that could diagnose different types of dementia in just minutes has been developed by Southampton researchers.
-

Finding cancer treatments using AI
Southampton researchers are using AI to study how immune cells function in the body, aiming to improve immunotherapy treatments and patient outcomes.
-

Insights from Mars transforming healthcare technology
Laser and light-based techniques lie behind both the search for life on Mars and accelerated drug discovery on Earth.
-

Transforming early cancer diagnosis
University of Southampton researchers are developing a new diagnostic blood test that uses biomarkers and artificial intelligence to predict multiple cancers at an early stage.
-

Adapting to warmer waters
Southampton PhD student Karolina Zarzyczny is improving our understanding of the mass movement of marine life in our oceans due to global heating.
-

Southampton researchers develop technologies to make better artificial limbs for people in Cambodia
Data science and, computer-aided design informed by local needs and skills help academic team to do prosthetics differently.
-

Encouraging healthier eating for adolescents
University researchers have conducted focus groups with young people to understand the different factors that play a role in their food choices.
-

Improving health and wellbeing using molecular imaging
Innovative molecular imaging by Southampton researchers is shedding light on the interaction of pharmaceutical drugs with the gut microbiome and the repercussions this has on health and wellbeing.
-

Understanding how best to treat ADHD: a dual approach
Extensive Southampton-led research that explored the best way to treat attention deficit hyperactivity disorder (ADHD) is having international impact.
-

Supercharging your immune system
Research led by Professor Nullin Divecha and his laboratory team has identified a way to boost the power of the human immune system to find and destroy cancer cells within the body.
-

Using copper to fight COVID-19
Biological Sciences and Health Sciences are showing how copper can provide a permanent defence against the spread of Coronavirus on surfaces in just one minute.
-

Helping diagnose and treat COVID-19 more quickly
When COVID-19 hit in late 2019, a worldwide fight to beat it began. Our biomedics have taken on the challenge to diagnose it quicker and treat it better - with promising early successes.
-

Empowering young people to make positive health choices during the COVID-19 pandemic
Helping young people to make positive lifestyle choices has never been as important as it is now. LifeLab is working to ensure young people have a voice, and that they understand the choices they can make to reduce the impact COVID-19 on their lives.
-

Using nanoclay gel to regrow, repair and replace damaged cells
Southampton researchers have developed an innovative medical clay that can be used to apply regenerative medicine to patients with musculoskeletal conditions.
Related research institutes, centres and groups
-

Adolescent Health and Wellbeing
We focus on the physical and mental health and well-being of young people. We aim to ensure healthy futures for them and future generations. -
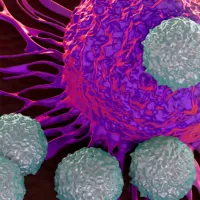
AI for the Life Sciences
Modern biology increasingly uses the generation and interpretation of large datasets to understand the complex processes that underpin life, health and disease. AI gives us tools and techniques to do this. -

BiOmics
Technological advances have allowed scientists to gather large amounts of data about a vast array of species, organisms and single cells. Our researchers are using mathematical modelling, machine learning and other algorithms to extract information and patterns from large data sets to further our understanding of disease. -
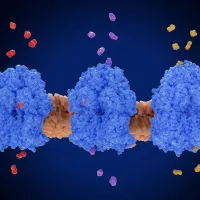
ChemLife Network
Advances in chemistry and molecular biology have revolutionised our understanding of life’s building blocks. We design, explore, and apply these molecules to advance biotechnology, medicine, and sustainability. -
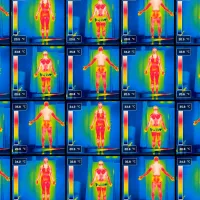
Climate Change Health and Wellbeing
Climate action, reducing emissions and adapting to change, protects our health and future wellbeing, especially when we highlight and support the health benefits these actions bring. -

Digital Health
Our researchers are examining and developing information and communication technologies to help address the health problems and challenges faced by patients. -
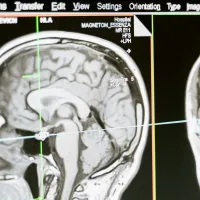
Interdisciplinary Dementia and Ageing Centre
We are a network that brings together local expertise from across the research spectrum within the University and University Hospital Southampton. Our aim is to increase understanding of dementia and brain ageing and how to treat and care for people with these conditions. -
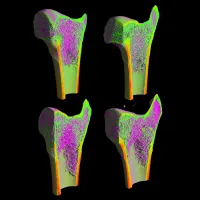
Interdisciplinary Musculoskeletal Health
The University hosts a substantial interdisciplinary community of researchers working to transform musculoskeletal health across the life course. -

Microfluidics and Sensors
Microfluidics is the interdisciplinary study of the behaviour, manipulation and application of fluid at the microscale. It underpins the concept of the lab-on-a-chip, where multiple key components and operations are integrated onto one small platform. -

National Biofilms Innovation Centre (NBIC)
NBIC is the central hub where academia, industry, government and public policy come together to tackle the grand challenges biofilms present, impacting $4 trillion in global economic activity. -

Southampton Imaging
Imaging has become an essential part of scientific research, from biomedical sciences to engineering to optoelectronics.
Connect with us
Contact us
Life Sciences Building (85), Highfield Campus, University of Southampton, SO17 1BJ
View in Google Maps